You are here
Deconstructing SMA: from RNA Processing to Motor Neuron Disease
Speakers
Abstract
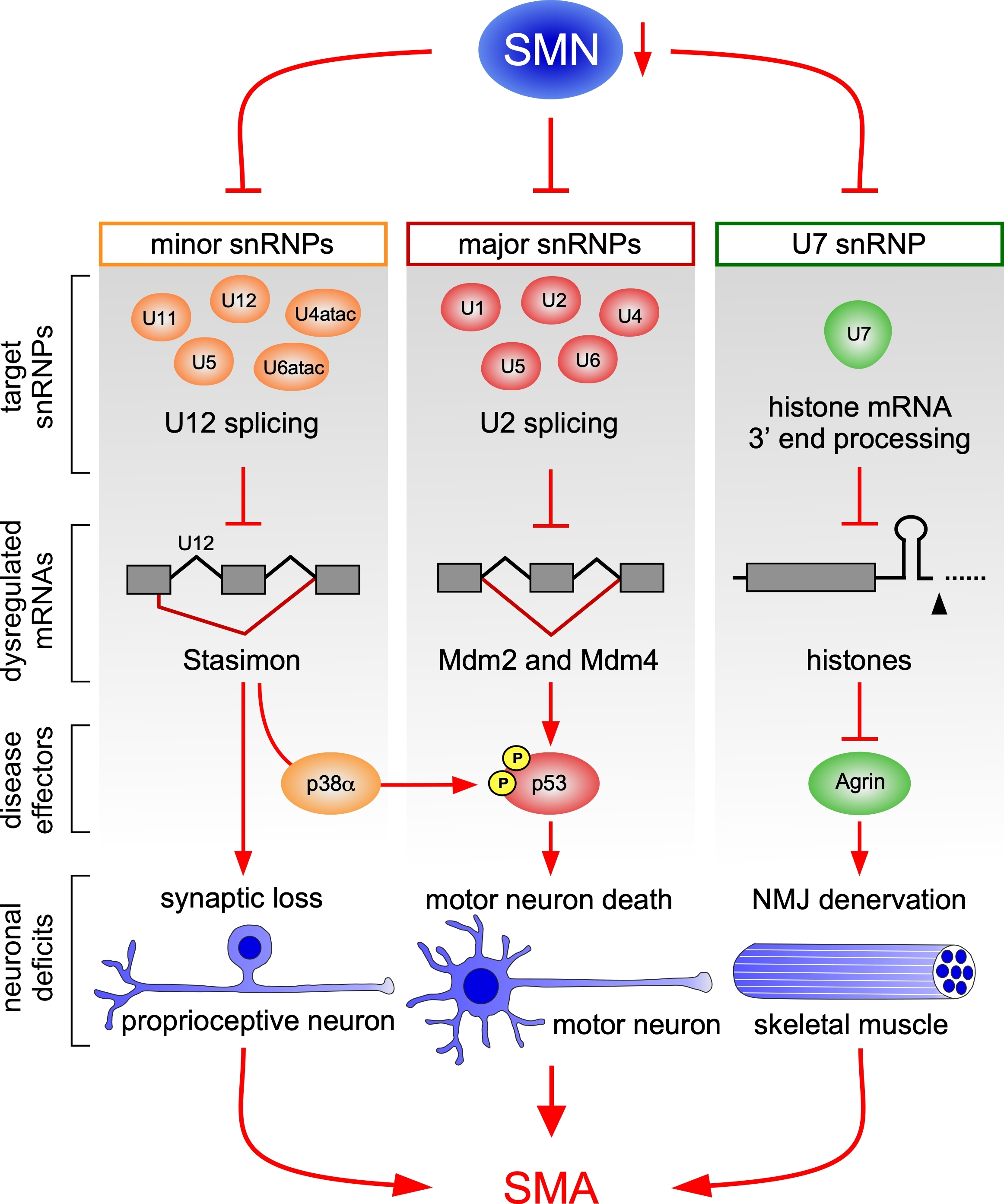
Research in the Pellizzoni laboratory investigates the mechanisms by which RNA binding proteins (RBPs) and molecular chaperones that mediate their assembly into RNAprotein complexes (RNPs) regulate gene expression. The laboratory also focuses on the question of how general perturbations of RNA metabolism cause synaptic dysfunction and neuronal death in neurodegenerative diseases such as spinal muscular atrophy (SMA). In these studies, we employ cellular and animal models as well as a wide range of genomic, biochemical, molecular, and imaging approaches. High-throughput screens are also used to discover chemical and genetic modifiers of disease pathways. On one hand, these efforts are designed to advance our knowledge of how RNA regulation contributes to neural circuit function. On the other, they aim to deconstruct disease mechanisms and identify potential therapeutic targets.